Discoveries in DNA: What's New Since You Went to High School?
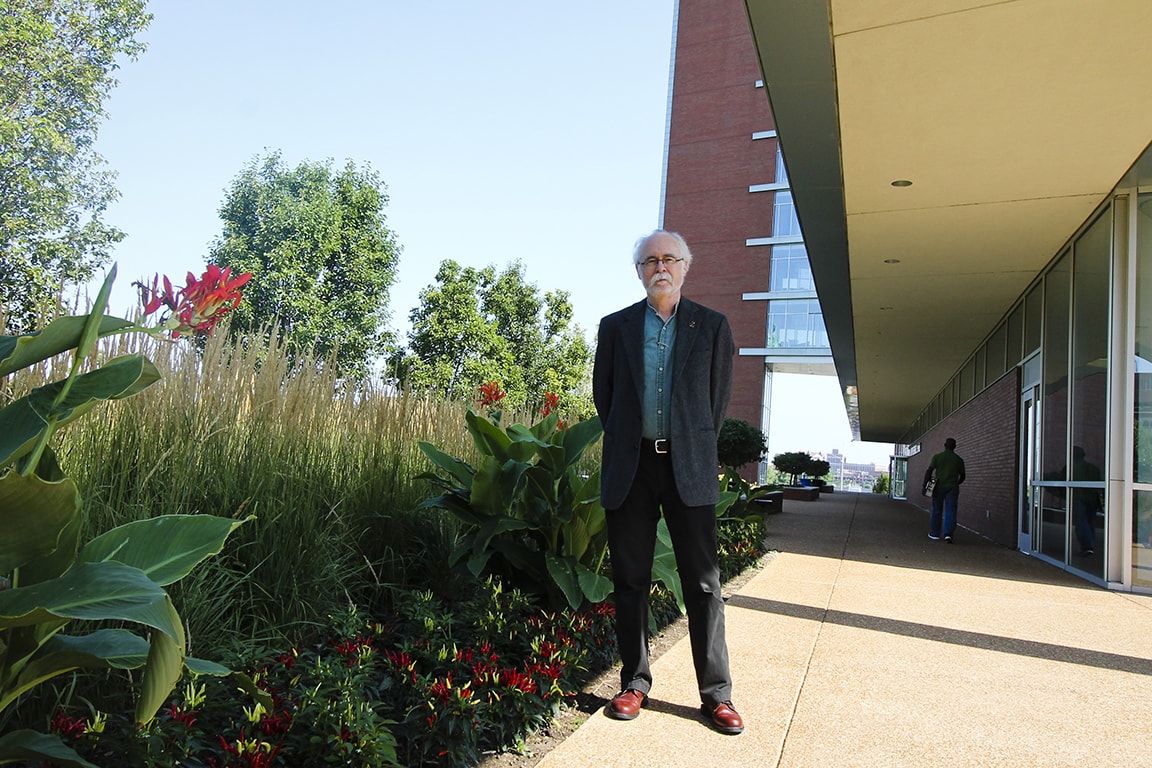
Joel Eissenberg, Ph.D., associate dean for research and professor of biochemistry and molecular biology at Saint Louis University School of Medicine, shares highlights from recent decades in the field of genetics.
Scientific and technological advances in the last 50 years have led to extraordinary progress in the field of genetics, with the sequencing of the human genome as both a high point and starting point for more breakthroughs to come.
Advances in molecular genetics have propelled progress in fields that deal with inherited diseases, cancer, personalized medicine, genetic counseling, the microbiome, diagnosis and discovery of viruses, taxonomy of species, genealogy, forensic science, epigenetics, junk DNA, gene therapy and gene editing.
For many non-scientists, a recap may be in order. With such rapid progress, the field has moved well beyond the knowledge covered in many biology classes over the years.
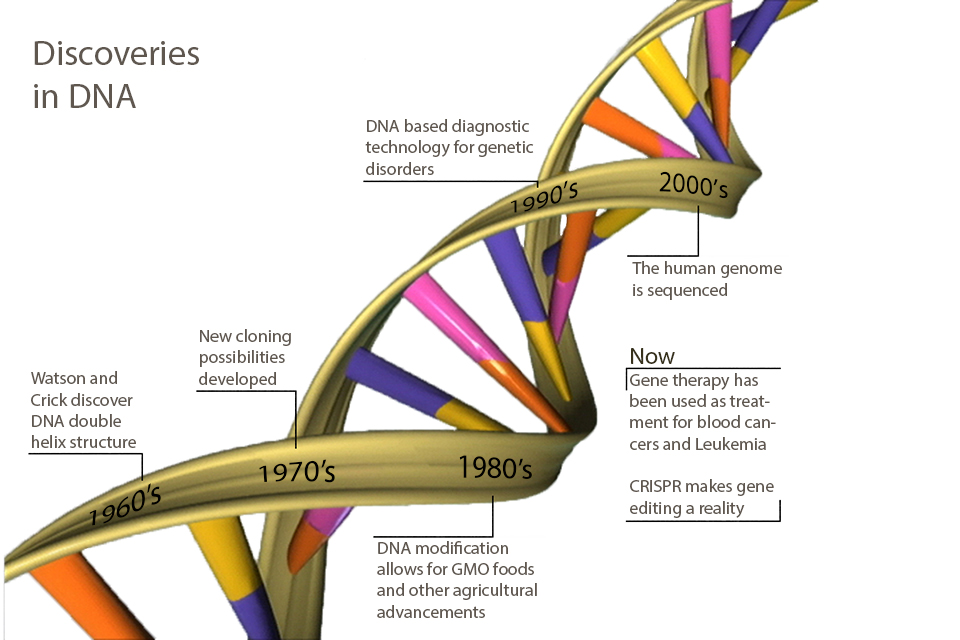
If you took high school biology in the 1960s, you probably learned about DNA as the genetic material and the structure of the DNA double helix (published in 1953 by Watson and Crick). You may also have learned about the genetic code, by which the sequences of DNA encode amino acids (worked out by Nirenberg, Khorana and colleagues by 1961).
If you took high school biology in the 1970s, you probably also learned about cloning (worked out by Herb Boyer, Stanley Cohen and Paul Berg by 1972) and the potential for recombinant DNA technology to provide gene therapy, create novel drugs and improve agriculture.
If you took high school biology in the 1980s, you may have learned about the clinical use of recombinant human insulin for diabetes treatment (approved for the Eli Lilly products in the US by the FDA in 1982). In agriculture, the use of Agrobacterium tumefacians as a bacteria-mediated delivery system to transfer recombinant DNA to crops (developed by Mary-Dell Chilton and colleagues in the 1970s) marked the advent of GMO foods and other commercial plant products.
If you took high school biology in the 1990s, you probably learned about the molecular basis for human genetic disorders such as cystic fibrosis (1989), Huntingtons (1993), Duchenne and Becker muscular dystrophy (1987), and a rapidly growing list of single-gene disorders, and the correspondingly rapid growth in clinical diagnostic technology based on DNA sequence information, enabling certain diagnosis, sometimes before the advent of overt symptoms.
If you took high school biology in the ‘00s, you probably heard about the completion of the human genome sequence. The completion of a “rough draft” was announced by President Bill Clinton and British Prime Minister Tony Blair in 2000, although a more-or-less complete sequence was only finalized in 2006. You may also have learned that this achievement heralded the arrival of the age of personalized medicine.
The big breakthrough in decoding the human genome was the invention of technology to obtain large amounts of DNA sequence, which began in the 1970s with the work of Ray Wu, Walter Gilbert, Fred Sanger and their colleagues to establish the core strategies for obtaining continuous sequence information for DNA chains. This included advances in recombinant DNA technology—for example, the creation of recombinant artificial chromosomes—combined with semi-automated (invented in the Leroy Hood lab in 1986) and later automated DNA sequencing. Today, the goal of obtaining the complete sequence of an individual genome for $1,000 is nearly within reach.
Although it isn’t biology, it must be acknowledged that the human genome sequencing
project also required parallel advances in computer speed and storage to acquire,
store and manipulate billions of nucleotides of DNA sequence. The assembly and analysis
of human tumor cell genomes, many of which contain chromosome deletions, duplications
and insertions, as well as single nucleotide changes, requires immense data storage
capacity and high-speed computation.
The invention of the Polymerase Chain Reaction (PCR) technology by Kary Mullis and
colleagues in 1985 transformed molecular genetics. This had immediate application
for DNA diagnostics, because once a gene implicated in an inherited disorder has been
identified and sequenced in its normal form, PCR could be used to amplify the corresponding
sequences from patient DNA in a matter of hours, with sequencing of the PCR products
to identify the exact molecular mutation in a matter of days at that time, and today
in a matter of hours. PCR has also seen application in the identification of emerging
pathogens.
The findings made by scientists during these decades have led to advances in many
different fields. Explore the following topics to learn more about how breakthroughs
in molecular genetics are being applied in the world.
Thus far, the impact of molecular genetics on human disease has been primarily to identify specifically which genes are implicated in specific diseases. For single-gene disorders, mutations have been discovered in hundreds of genes. The current challenge is to identify which genes contribute to multifactorial conditions like obesity, heart disease, alcohol dependency, schizophrenia and autism. So far, “genome-wide association studies” have identified variant DNA sequences showing statistical association with these and other complex diseases, but demonstrating a mechanistic role for these variants has proven elusive.
Rapid and inexpensive genome sequencing, together with high-speed informatics and a large and expanding database of annotated human DNA sequence variants associated with disease risk, has made possible personalized medicine and customized genetic counseling. In a particularly famous case, actress Angelina Jolie elected to have a double mastectomy and her ovaries and fallopian tubes surgically removed when she learned that she carried a mutation in the BRCA1 gene that predicted an 87 percent risk of breast cancer and a 50 percent risk of ovarian cancer. Unfortunately, for the vast majority of genetic cancer associations, removing nonessential tissue is not an option. However, prior knowledge of an increased risk can result in increased surveillance, and cancer is most curable when caught early. For example, persons at increased risk for hereditary non-polyposis colorectal cancer should undergo frequent colonoscopies to identify and remove pre-cancerous colon polyps before they turn into full-blown colon cancer. Since 23andMe began offering direct-to-consumer marketing of genomic sequencing, over a dozen companies now offer various forms of genome sequencing and analysis.
A healthy intestine carries about 10 times as many microorganisms as the number of cells in the entire body. The metabolic activity of these microorganisms can significantly impact health. For example, their metabolic activity is an important source of biotin (vitamin B7), and the composition of gut microflora can shape the immune response, leading to sensitivity or resistance to allergies and autoimmunity. The availability of a large number of complete microbial genomes and the technology of high-volume DNA sequencing has enabled the genotyping of gut microbiomes under different dietary and health conditions, leading to new, detailed understanding of the differences between healthy and unhealthy gut microflora. Direct-to-consumer gut microflora sequencing services are currently available, although the benefits of this knowledge for otherwise healthy people are currently limited.
Most high school biology students learn some basic animal and plant taxonomy. The foundation of classical taxonomy is morphology. With the availability of genome sequences from representative species in whole phyla, rigorous quantitative measurements of genetic distance based on DNA sequence divergence has been used to test existing evolutionary trees and to re-classify organisms in all kingdoms. For example, large-scale taxonomic DNA sequence comparisons have established more rigorous relationship trees and taxonomic distances for the large and diverse class Aves (birds) and the phylum Arthropoda.
Human genome sequencing provides much more detailed and specific genealogical information. Several commercial services will provide information on likely ancestry based on combinations of DNA sequence variants known to be rare or prevalent among people originating from different regions of the world. However, it should be noted that one sometimes unwelcome outcome of genome sequences for pedigree or genealogical purposes is the discovery of non-paternity. While rates vary widely between different populations, they have ranged between 2 and 30 percent in specific studies.
Forensic science is increasingly turning to DNA sequencing to implicate or exonerate potential culprits and to identify remains. In such cases, it is usually sufficient to sequence only a subset of genomic DNA representing regions found to be most variable among individuals. This approach circumvents the much higher cost of sequencing and data management for whole genome sequencing, while providing sufficient specificity for forensic purposes.
Biochemists have known that human DNA (as well as the DNAs of many microbes, plants and animals) contains other bases besides the canonical adenine, cytosine, guanine and thymine (ACGT). In human chromosomes, 3 to 5 percent of the cytosine bases are actually a modified form of cytosine called 5-methyl cytosine. This modification is usually associated with repression of genes in humans. Importantly, the extent of modifications can differ at the same gene in the sperm and egg, such that the expression of dad’s copy will be different from mom’s copy in the child that inherits them. This phenomenon is called “imprinting.” Parental imprinting is important for genetic health, as failure of this imprinting underlies syndromes such as Prader-Willi, Angelmans, Beckwith-Weidemann and Silver-Russell syndromes.
The transmission of different states of gene expression through multiple cell divisions and across generations has been termed “epigenetic,” since the underlying DNA sequence is identical in both states. While the four bases of DNA – adenine, cytosine, guanine and thymine – cannot be altered by a parent’s life experiences, scientists have discovered that a form of one of the base pairs, cytosine, can be expressed in different forms as a result of environmental factors. Techniques have been developed over the last decade to allow for “epigenomics,” the genome-wide characterization of methylation patterns. Epigenomics is presently being used to identify inherited pre-disposition to obesity, diabetes, cardiovascular disease, addiction and psychiatric disorders, as well as markers for aging and cancer progression.
Drugs that inhibit or stabilize epigenetic marks are in clinical use for cancer and are being tested for other indications like sickle cell disease.
In the past 15 years, however, detailed analysis of which regions of DNA are transcribed into RNA copies has uncovered a significant amount of non-protein-coding RNAs that serve regulatory functions. So-called microRNAs have been shown to target specific protein-coding RNAs for destruction or inhibition of protein synthesis. Other RNAs, termed long noncoding RNAs, also appear to regulate expression of protein-coding RNAs, but the mechanisms are just now being worked out. That said, this still leaves a majority of our genome with no apparent function. It seems likely that the mechanisms by which DNA accumulates in genomes through transposon jumping and genome duplications is not balanced with a similar rate of sequence elimination, and that the burden of this unpurged DNA has little or no evolutionary cost.
A major goal for gene sequencing has always been as a platform for the design and implementation of gene therapy. The first gene therapy was bone marrow engraftment for the treatment of leukemia and other blood cancers. The first human bone marrow transplant was performed in 1956 by E. Donnall Thomas. In these therapies, the patient’s own blood-producing bone marrow cells are treated with radiation and chemotherapy, and the blood of a twin or closely matched donor is instilled. The idea is that the ablation will destroy any remnant of the cancer together with the healthy hematopoietic stem cells, and that donor’s blood stem cells will populate the patient’s marrow and regenerate the entire red and white blood cell repertoire from healthy cells. In effect, this is gene therapy, since the donor’s genes are replacing the patients genes in the blood cell lineages.
Targeted gene therapies, however, had to wait for (1) the identification of the genes to target, (2) the cloning and/or sequencing of the relevant genes and in some cases, the specific disease-causing variant, (3) a full understanding of the normal gene function and regulation, and (4) the development of efficient ways to deliver genes to the relevant tissues at therapeutic levels. The advent of molecular cloning, DNA sequencing and the many tools of molecular genetics and cell biology has given us sufficient knowledge of the basis for disease and the genes to target, but what has limited the application of gene therapy has been efficient gene delivery systems.
Scientists realized viruses could be the perfect tool to do the work of gene editing.
They are already designed by nature to insert themselves into our DNA. The first successful
targeted human gene therapy was reported in 2000. This was a virus-mediated therapeutic
gene to treat X-linked severe combined immunodeficiency. The deck was stacked in favor
of success in the choice of this particular disease since it was known that the therapeutic
gene just had to be expressed in a modest number of blood cells to achieve therapeutic
benefit. The therapeutic gene was carried by an engineered retrovirus that was used
to infect the patient’s blood cells before they were re-injected into the patient.
Thus far, targeted gene therapy successes have been very limited. Other blood disorders
that have shown significant benefit from targeted gene therapy in small trials include
hemophilia (specifically, factor IX deficiency), severe beta-thalassemia (deficiency
for the adult beta-globin gene) and leukemia, where the patient’s immune cells were
treated to enable them to recognize cancer cells and destroy them. Targeted gene therapy
for degenerative blindness caused by Leber congenital amaurosis improved vision for
a few years but failed to arrest the degeneration process.
The first approved commercial targeted gene therapy is Alipogene tiparovovec (trade name Glybera), a virus-mediated therapeutic delivery of human lipoprotein lipase to the muscle cells of lipoprotein lipase deficiency patients. It was approved for clinical use in Europe in 2012.
For many genetic disorders, the disease results from the expression of a defective gene product, not the complete absence of the product. For example, sickle cell disease and the major form of cystic fibrosis are both associated with abnormal proteins. In such cases, editing a patient’s own gene to the normal form should provide greater benefit than merely expressing the normal protein in the presence of the abnormal one.
For gene editing to work, it is essential to uniquely target a single site among the
3 billion nucleotides in the haploid (single set of unpaired chromosomes) human genome.
In other words, therapeutic gene editing therapy must be able to efficiently edit
the intended target without introducing off-target editing at sites that fortuitously
resemble the intended target.
Two targeting strategies are currently under development—protein-based targeting and
RNA-based targeting. In both cases, the idea is to target an enzyme that cuts both
strands of the double helix at a specific site. If the goal is to inactivate the target
gene, the creation of the break is sufficient to trigger cellular mechanisms that
lead to error-prone repair and inactivating mutations. True editing—the replacement
of bad sequence with good sequence—requires the simultaneous introduction of DNA fragments
containing good sequence into the same cells.
Protein-based targeting strategies rely on custom-engineered modular proteins that
recognize and bind to specific DNA sequences. One approach builds on the so-called
zinc finger protein fold first described in sequence-specific transcription factors.
The publicly traded biotech company Sangamo BioSciences was founded in 1995 to exploit
zinc finger protein engineering for gene therapy and agricultural genetic engineering.
In this approach, a series of zinc finger modules, each chosen to recognize a specific
3-nucleotide motif, are fused in tandem to one another and to a nuclease subunit.
Another protein-based targeting strategy, TALENs, is based on the transcription activator-like
effector (proteins secreted by Xanthomonas bacteria). In this case, repeating 33-34
amino acid modules with specificity to each of the four bases in DNA are fused in
tandem to create the targeting peptide.
More recently, the clustered regularly-interspaced short palindromic repeats (CRISPR)
prokaryotic immunity mechanism has been exploited for gene editing. In this case, specific DNA sequences are targeted by RNA-DNA hybridization, which
directs the Cas9 enzyme to cleave DNA at the target site. This mechanism was only
worked out in 2007, but has already emerged as the front-runner technology for gene
editing, due to its relative simplicity and high efficiency. In human cells, the efficiency
of zinc-finger- and TALE-mediated editing achieve efficiencies of 1 to 50 percent,
while CRISPR-Cas9 editing has been reported to have efficiencies of up to 78 percent
in single-cell mouse embryos. Exciting clinical applications of gene editing include
correcting the mutation in the bone marrow stem cells of patients with sickle cell
disease or hemophilia.
One of the most exotic applications of genetic engineering to be proposed is the resurrection of woolly mammoths. A project underway at Harvard University, under the direction of geneticist Dr. George Church, seeks to edit the elephant genome to create a cold-tolerant mammoth-like chimera using CRISPR/Cas9 technology. This strategy is enabled by the complete sequencing of the mammoth and elephant genomes, and based on gene annotations that suggest the target genes most likely to program the mammoth adaptations to extreme cold into elephant DNA. The stated goal is to repopulate the tundra and boreal forest in Eurasia and North America, to protect endangered Asian elephants and to revive an ancient grassland in the tundra, with the hope of forestalling the melting of Siberian permafrost.